More Information
Submitted: March 30, 2026 | Approved: April 10, 2026 | Published: April 13, 2026
Citation: Elafari MA, Bibat MA, Ayoub M, Slaoui A, Karmouni T, Koutani A, et al. The Urinary Microbiome: Shifting Paradigms from Sterile Urine to Microbial Dysbiosis in Chronic Pelvic Pain Syndrome. Int J Clin Microbiol Biochem Technol. 2026;9(1): 022-026. Available from:
https://dx.doi.org/10.29328/journal.ijcmbt.1001036
DOI: 10.29328/journal.ijcmbt.1001036
Copyright license: © 2026 Elafari MA, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Keywords: Urinary microbiome; Urobiome; Chronic pelvic pain syndrome; IC/BPS; CP/CPPS; Dysbiosis; Metagenomics; Urological disorders
The Urinary Microbiome: Shifting Paradigms from Sterile Urine to Microbial Dysbiosis in Chronic Pelvic Pain Syndrome
Mohammed Amine Elafari* , Mohammed Amine Bibat, Mamad Ayoub, Amine Slaoui, Tarik Karmouni, Abdelatif Koutani and Khalid Elkhader
, Mohammed Amine Bibat, Mamad Ayoub, Amine Slaoui, Tarik Karmouni, Abdelatif Koutani and Khalid Elkhader
Urology B Department, IBN SINA Hospital, University Hospital Center IBN SINA, University Mohammed V, Rabat, Morocco
*Corresponding author: Mohammed Amine Elafari, Urology B Department, Ibn Sina Hospital, University Hospital Center IBN SINA, University Mohammed V, Rabat, Morocco, Email: [email protected]
The prevailing dogma regarding urinary tract sterility has been fundamentally challenged by advances in culture-independent molecular techniques. The urinary microbiome, also known as the urobiome, is defined as a complex ecosystem comprising bacteria, fungi, and viruses. This microbiome plays a pivotal role in maintaining urological health. Emerging evidence suggests a potential link between urinary microbial dysbiosis and the pathogenesis of Urologic Chronic Pelvic Pain Syndrome (UCPPS), a clinical umbrella term that explicitly encompasses two major phenotypes: Interstitial Cystitis/Bladder Pain Syndrome (IC/BPS) and Chronic Prostatitis/Chronic Pelvic Pain Syndrome (CP/CPPS).
This review examines the paradigm shift in understanding urinary tract microbiology, characterizes the urobiome in health and disease, and explores the therapeutic implications of microbiome-targeted interventions for UCPPS management.
For over a century, the urinary tract was regarded as a sterile environment in healthy individuals, with bacterial presence being interpreted as pathological contamination or infection [1-3]. This paradigm was primarily based on standard urine culture techniques, which detect only a limited spectrum of cultivable organisms under specific laboratory conditions [4,5]. The advent of Next-Generation Sequencing (NGS) technologies, particularly 16S rRNA gene sequencing and metagenomic approaches, has revolutionized our understanding of urinary microbiology by enabling detection of low-biomass microbial communities that evade conventional culture methods [6-8].
Throughout this review, UCPPS is discussed as a unified clinical entity, while specific microbial distinctions are highlighted for its primary phenotypes, IC/BPS and CP/CPPS, to provide a comprehensive understanding of the syndrome’s heterogeneous nature.
The urinary microbiome, termed the “urobiome,” comprises diverse bacterial genera, along with fungal and viral components, that colonize the urinary tract in both healthy and diseased states [5,9]. This discovery has profound implications for understanding urologic chronic pelvic pain syndrome (UCPPS), a debilitating condition affecting millions worldwide that encompasses IC/BPS and CP/CPPS [10,11]. Despite extensive research, the etiology of UCPPS remains poorly understood, with limited therapeutic options available. A mounting body of evidence indicates that urinary microbial dysbiosis may play a role in the chronic inflammation and pain associated with these conditions [10,12,13].
The healthy urobiome: Composition and characteristics
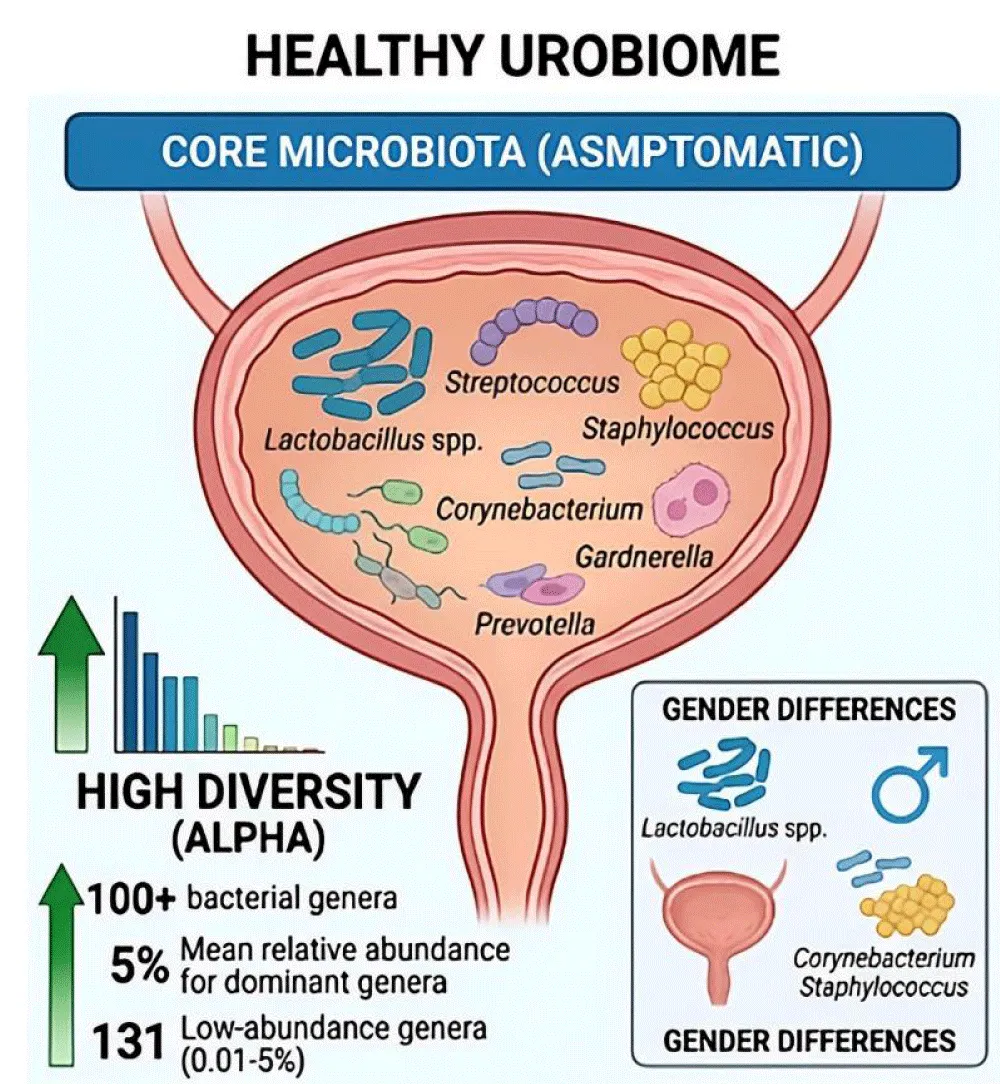
A healthy urobiome exhibits considerable diversity, with studies identifying over 100 bacterial genera in urine from asymptomatic individuals [5,14]. The core microbiota includes dominant genera such as Lactobacillus, Streptococcus, Staphylococcus, Corynebacterium, Gardnerella, and Prevotella, with mean relative abundances exceeding 5% [14,15]. Additionally, 131 low-abundance genera (0.01% – 5% relative abundance) contribute to the overall microbial ecosystem [14] (Figure 1).
Figure 1: Conceptual overview of the healthy urinary microbiome (urobiome). The healthy state is characterized by high alpha-diversity, featuring over 100 bacterial genera. The core microbiota is dominated by Lactobacillus, Streptococcus, and Corynebacterium. Significant gender-specific differences exist: females exhibit a Lactobacillus-rich profile providing protective functions, while males show a higher prevalence of Corynebacterium and Staphylococcus.
Significant gender-related differences characterize the urobiome. The urinary microbiota of female subjects characteristically exhibits elevated levels of Lactobacillus species, which have been demonstrated to offer protective functions against uropathogen colonization [4,5,14]. In contrast, male urobiomes demonstrate a greater prevalence of Corynebacterium, Staphylococcus, and other genera, a phenomenon that is likely attributable to anatomical and hormonal differences [5,14,15]. Furthermore, age exerts a significant influence on urobiome composition, with distinct microbial profiles being observed across various age groups [3,14].
Methodological considerations play a pivotal role in the characterization of urobiomes. Studies employing expanded quantitative urine culture (EQUC) and 16S rRNA sequencing have revealed that the sampling method affects the detected microbial diversity, with first-catch urine, midstream urine, and catheterized specimens yielding different results [6,15]. These findings underscore the importance of standardized collection protocols in urobiome research.
Urinary dysbiosis in chronic pelvic pain syndrome
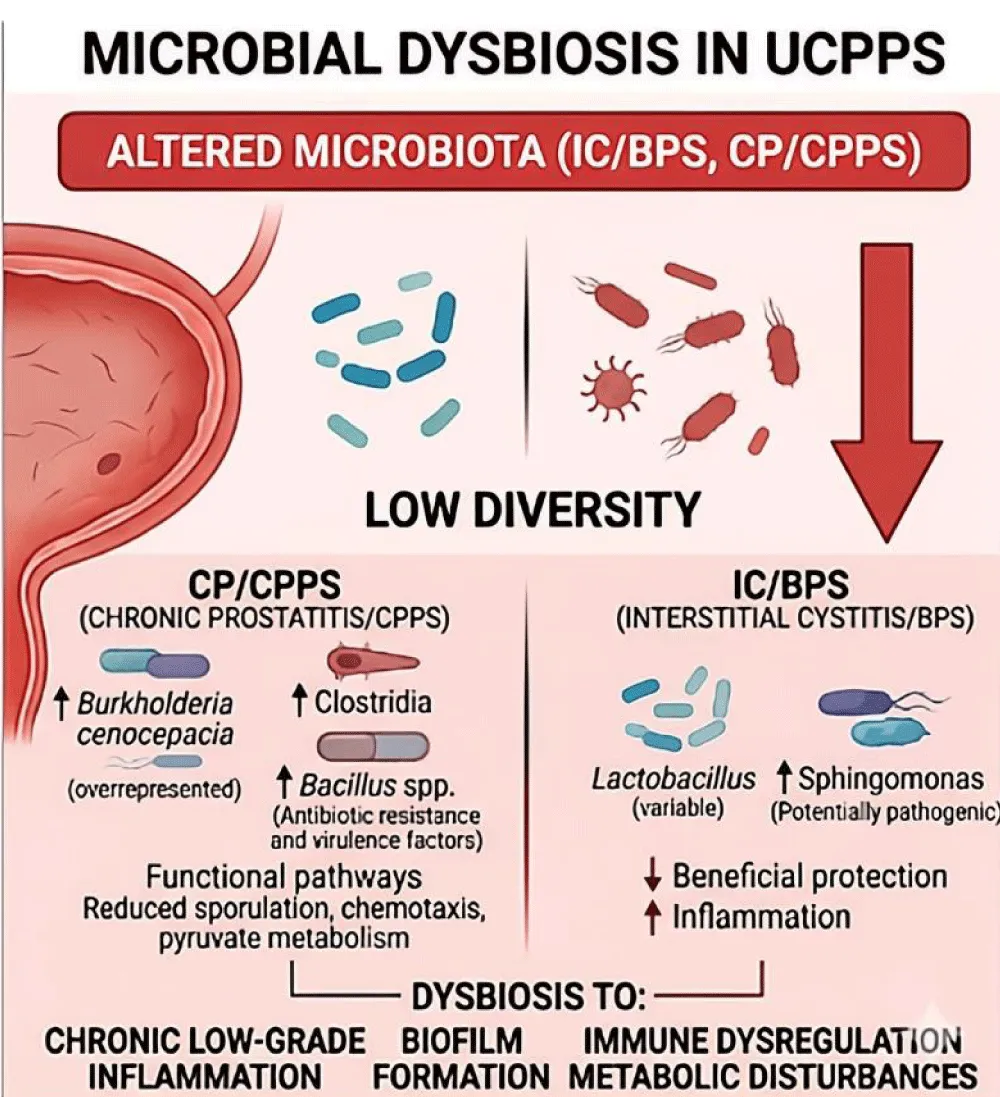
Microbial alterations in CP/CPPS: A multitude of studies have documented substantial disparities in urinary microbiome composition between patients with CP/CPPS and healthy controls. A seminal study by Shoskes, et al. demonstrated that patients with chronic prostatitis/chronic pelvic pain syndrome (CP/CPPS) exhibited significantly higher alpha-diversity (phylogenetic diversity) compared to control subjects, with distinct clustering patterns on principal coordinate analysis [12]. Specifically, patients with CP/CPPS showed overrepresentation of 17 bacterial clades, including Clostridia, and underrepresentation of 5 clades, notably Bacilli [12]. These compositional changes were associated with predicted perturbations in functional metabolic pathways, including sporulation, chemotaxis, and pyruvate metabolism [12] (Figure 2).
Figure 2: Microbial Signatures and Functional Implications of Dysbiosis in UCPPS. Distinct taxonomic shifts characterize the two main phenotypes: CP/CPPS shows enrichment of Clostridia and Burkholderia cenocepacia alongside increased virulence factors; IC/BPS is associated with fluctuations in Lactobacillus and enrichment of Sphingomonas. These shifts correlate with altered metabolic pathways and pro-inflammatory signaling that contribute to pain sensitization.
The Multidisciplinary Approach to the Study of Chronic Pelvic Pain (MAPP) Research Network identified Burkholderia cenocepacia as significantly overrepresented in male patients with urinary tract pain (UCPPS) compared to control subjects in initial stream urine specimens [16]. More recent metagenomic analyses have identified enrichment of Bacillus species in patients with CP/CPPS, with associated antibiotic resistance genes and virulence factors that may contribute to the presence of localized inflammation, asss documented in the relevant literature [17].
Of particular significance is the correlation between microbiome differences and clinical phenotypes. Research employing the UPOINT classification system has identified distinct microbial signatures associated with specific symptom domains, including neurological and systemic features [12]. This finding suggests that urobiome profiling may eventually enable personalized therapeutic approaches based on individual microbial and clinical characteristics.
Microbial alterations in IC/BPS: A review of the extant literature on IC/BPS microbiome studies reveals a consistent finding of significant variations in urinary microbial diversity and species composition when compared to healthy controls [11]. The most prominent finding involves alterations in Lactobacillus abundance, though the direction of change varies across studies [11,18]. Some investigations report decreased Lactobacillus in IC/BPS patients, potentially compromising protective functions against pathogen colonization and inflammatory responses [11].
Integrated microbiome-metabolome analyses have revealed that microbial dysbiosis in IC/BPS correlates with altered urinary metabolite profiles [18]. Differential genera such as Lactobacillus and Sphingomonas show associations with specific metabolites, suggesting that microbe-metabolite interactions may influence the inflammatory microenvironment [18]. Lactobacillus appears to exert protective effects, while Sphingomonas may represent a pathogenic factor [18]. The metabolite theophylline, identified as differentially abundant in IC/BPS patients, It has been demonstrated that the subject exhibits anti-inflammatory properties that may modulate disease activity [18]. Gender-specific differences in the urinary microbiome of patients with interstitial cystitis/bladder pain syndrome (IC/BPS) mirror those observed in healthy populations. Specifically, female patients exhibit higher levels of Lactobacillus and Escherichia-Shigella compared to males, while males exhibit greater abundance of Bacteroides and Acinetobacter [18].
Symptom flares and fungal dysbiosis: A study by the MAPP Network examined symptom flares in female UCPPS patients and revealed a striking association between fungal presence and acute symptom exacerbations [19]. The study’s participants who reported symptom flares demonstrated significantly greater prevalence of fungi, particularly Candida and Saccharomyces species, in midstream urine (15.7%) compared to those without flares (3.9%) [19]. This association persisted after adjusting for antibiotic use and menstrual phase, with an odds ratio of 8.3 for fungal presence during flares [19]. These findings suggest that fungal dysbiosis may trigger or perpetuate symptom exacerbations in susceptible individuals.
Gut-bladder axis: The field has recently seen a proliferation of evidence indicating that gut microbial dysbiosis may play a role in the pathogenesis of UCPPS [10]. Systematic reviews have identified reduced diversity and altered composition of intestinal microbiota in UCPPS patients compared to controls [10]. This suggests the existence of a potential gut-bladder axis, whereby intestinal dysbiosis influences urinary tract inflammation through systemic immune modulation, bacterial translocation, or metabolite-mediated mechanisms [10,13].
Methodological advances in urobiome research
The advent of several technological advances has facilitated the detection and characterization of the urobiome. The most widely employed method remains 16S rRNA gene sequencing, which provides taxonomic identification of bacterial communities with high sensitivity [2,6,7]. However, this approach has limitations, including the inability to detect non-bacterial microorganisms and limited species-level resolution [8].
Metagenomic sequencing offers more comprehensive characterization, enabling identification of bacteria, fungi, viruses, and functional gene content, including antibiotic resistance genes and virulence factors [8, 17]. Studies employing shotgun metagenomics have revealed the presence of diverse eukarya (median 2 species/sample) and viruses (median 3 viruses/sample) in addition to bacterial communities (median 41 genera/sample) [8].
EQUC represents a culture-based approach that employs extended incubation periods, diverse culture media, and aerobic/anaerobic conditions to detect fastidious organisms missed by standard urine culture [6,7]. When combined with molecular methods, it has been demonstrated that the provision of complementary information regarding cultivable microbiome members enables the execution of functional studies of isolated organisms [2,6]. However, a critical challenge for the field remains the standardization of methodologies, including DNA extraction protocols, sequencing platforms, and bioinformatic pipelines [1,6,9]. Variations in methodology contribute to heterogeneity across studies and limit the capacity for direct comparisons of findings.
Pathogenic mechanisms: From dysbiosis to disease
A multitude of mechanisms have been postulated to establish a link between urinary dysbiosis and the pathogenesis of UCPPS. Chronic low-grade inflammation is a hallmark of the condition, potentially driven by persistent microbial stimulation of urothelial and immune cells [1,9,13]. Specific bacterial species may produce pro-inflammatory metabolites, toxins, or virulence factors that activate inflammatory pathways and sensitize pain receptors [12,17].
Biofilm formation by urobiome members may create protected microbial communities resistant to host immune responses and antimicrobial therapies [1,9]. These biofilms can serve as persistent sources of inflammatory stimuli and may explain the chronic, relapsing nature of UCPPS.
Immune dysregulation, including autoimmune mechanisms, has been implicated in UCPPS, particularly CP/CPPS [13]. Microbial antigens may trigger aberrant immune responses in genetically susceptible individuals, leading to chronic inflammation even in the absence of active infection [13]. Molecular mimicry between microbial and host antigens could potentially drive autoimmune reactions targeting urinary tract tissues.
Metabolic disturbances resulting from dysbiosis have the potential to alter the urinary microenvironment, affecting pH, nutrient availability, and metabolite composition [12,18]. These changes can perpetuate dysbiosis and contribute to urothelial dysfunction and pain sensitization.
Therapeutic implications and future directions
The recognition of urinary dysbiosis in UCPPS has led to the emergence of novel therapeutic avenues. A variety of microbiome-targeted interventions are currently under investigation. These include probiotics, particularly Lactobacillus species, which are designed to restore a healthy composition to the microbiome [1,4]. However, clinical trials conducted to date have been underpowered and heterogeneous in terms of formulations, routes of administration, and outcomes, yielding inconclusive results [4]. Optimized probiotic preparations specifically designed for urinary tract colonization require further development and rigorous clinical testing.
Another potential approach involves targeted antimicrobial therapy based on urobiome profiling. The identification of specific pathogenic organisms or dysbiotic patterns could guide precision antibiotic selection [17]. Nevertheless, concerns regarding antibiotic resistance and the potential for further disruption of the urobiome necessitate a judicious approach to the implementation of this strategy.
Bacteriophage therapy offers a novel approach to selectively eliminating pathogenic bacteria while preserving beneficial microbiome members [1]. Phage preparations targeting specific uropathogens are under development, though clinical applications remain experimental.
Fecal microbiota transplantation (FMT) has demonstrated efficacy in treating recurrent Clostridioides difficile infection and has shown preliminary promise for UCPPS in small retrospective studies [4]. The gut-bladder axis provides a theoretical rationale for FMT in UCPPS; however, prospective trials are necessary to establish efficacy and safety.
Metabolite-based therapies that target specific dysbiosis-associated metabolic pathways represent an emerging frontier in this field [12,18]. The identification of protective metabolites, such as theophylline, or pathogenic metabolites, could enable the development of small-molecule interventions to modulate the urinary microenvironment.
The diagnostic applications of urobiome profiling have demonstrated considerable potential in the early detection of diseases, the stratification of phenotypes, and the prediction of treatment response [9,11]. Machine learning models that incorporate urobiome data have attained diagnostic accuracies surpassing 80% for specific urological conditions [9]. Analogous approaches have the potential to facilitate personalized UCPPS management based on individual microbial signatures.
The following table synthesizes the paradigm shift in urinary microbiology, illustrating the transition from the traditional sterility dogma to a contemporary understanding of the urobiome’s role in UCPPS pathogenesis (Table 1).
| Table 1: The Paradigm Shift in Urinary Microbiology and UCPPS Management. | |||
| Feature | Traditional View (Dogma) | Contemporary View (Urobiome) | Clinical Implication for UCPPS |
| Sterility | Urine is sterile in healthy individuals. | Complex ecosystem of bacteria, fungi, and viruses. | Dysbiosis, not just infection, drives symptoms. |
| Detection | Standard Urine Culture (detects limited organisms). | NGS (16S rRNA), Metagenomics, and EQUC. | Identification of "unculturable" pathogens or protective flora. |
| Core Flora | Pathological if present. | Lactobacillus (female), Corynebacterium (male). | Maintenance of health via protective niches. |
| Pathogenesis | Acute infection or unknown etiology. | Dysbiosis, Biofilms, Immune Dysregulation. | Chronic low-grade inflammation and pain sensitization. |
| Therapy | Broad-spectrum antibiotics. | Probiotics, Phage therapy, FMT, Metabolite modulation. | Precision medicine and restoration of microbial balance. |
Challenges and knowledge gaps
Notwithstanding the considerable progress that has been made, substantial challenges persist. Methodological heterogeneity across studies has been identified as a significant impediment to the execution of meta-analyses and the establishment of consensus [1,9,10]. Standardization of sample collection, processing, sequencing, and bioinformatic analysis is imperative for the advancement of the field.
The question of whether causality or correlation is the primary factor remains unresolved. While there is ample documentation of the associations between dysbiosis and UCPPS, further elucidation is necessary to determine whether microbial alterations are indeed primary pathogenic factors, secondary consequences of disease, or epiphenomena. This requires longitudinal studies and mechanistic investigations [1,9,10].
The low biomass nature of the urobiome poses technical challenges, including contamination risks from reagents, collection materials, and environmental sources [6,7]. Rigorous negative controls and contamination-aware bioinformatics approaches are essential for reliable results.
The dearth of longitudinal data, however, has impeded the development of a comprehensive understanding of the dynamics of the urobiome over time, during symptom flares, and in response to therapeutic interventions [1,9]. To elucidate the temporal relationships between dysbiosis and clinical outcomes, prospective cohort studies employing serial sampling methods are imperative.
The intricate interplay between the host and its microbiome, encompassing the influence of genetic factors on urinary microbiome composition and immune responses to urinary microbes, remains to be fully elucidated [5]. The integration of host genomics, transcriptomics, and immunological profiling with microbiome data holds promise in offering more comprehensive insights into the pathogenesis of UCPPS.
The paradigm shift from the conventional understanding of urine as sterile to the recognition of a complex urinary microbiome has transformed the field of urological health and disease. There is compelling evidence linking urinary dysbiosis to UCPPS, with distinct microbial signatures characterizing IC/BPS and CP/CPPS. Advancements in molecular technologies have facilitated comprehensive characterization of the urobiome and the identification of potential therapeutic targets. However, for these findings to be implemented in clinical practice, there is a need to address methodological challenges, establish causality, and conduct rigorous clinical trials of microbiome-targeted interventions. The urobiome signifies a promising frontier in the field of urology, with the potential to transform the diagnosis and treatment of chronic pelvic pain syndrome through personalized, microbiome-informed approaches.
Declaration
Ethical approval: Ethics approval has been obtained to proceed with the current study.
Ethical approval for this study (Ethical Committee N009-24) was provided by the Ethical Committee Ibn University Hospitals, Rabat Morocco on 22 January 2024.
Ethical approval (Revised): Ethical approval for this study was obtained from the Ethical Committee of Ibn Sina University Hospital, Rabat, Morocco (Approval No. N009-24; dated 22 January 2024).
Guarantor of submission: The corresponding author is the guarantor of submission.
Availability of data and materials: Supporting material is available if further analysis is needed.
Provenance and peer review: Not commissioned, externally peer-reviewed.
- Wei CR, Basharat Z, Osama M, Mah K, Waheed Y, Hassan SS. An overview of the association of the urinary tract microbiome with various diseases and implications for therapeutics. Mini Rev Med Chem. 2025:MRMC-EPUB-150499. Available from: https://doi.org/10.2174/0113895575398906250825113635
- Robino L, Navarro N, Canales-Huerta N, González MJ, Cruz E, Sauto R, et al. Urogenital microbiome, intracellular bacterial communities, and their contribution to urinary tract infections. Microbiol Spectr. 2025:e0124725. Available from: https://doi.org/10.1128/spectrum.01247-25
- Pastuszka A, Tobor S, Łoniewski I, Wierzbicka-Woś A, Sielatycka K, Styburski D, et al. Rewriting the urinary tract paradigm: the urobiome as a gatekeeper of host defense. Mol Biol Rep. 2025;52(1):497. Available from: https://doi.org/10.1007/s11033-025-10609-w
- Roth RS, Liden M, Huttner A. The urobiome in men and women: a clinical review. Clin Microbiol Infect. 2023;29(10):1242-1248. Available from: https://doi.org/10.1016/j.cmi.2022.08.010
- Brubaker L, Putonti C, Dong Q, Wolfe AJ. The human urobiome. Mamm Genome. 2021;32(4):232-238. Available from: https://doi.org/10.1007/s00335-021-09862-8
- Gasiorek M, Hsieh MH, Forster CS. Utility of DNA next-generation sequencing and expanded quantitative urine culture in diagnosis and management of chronic or persistent lower urinary tract symptoms. J Clin Microbiol. 2019;58(1):e00204-19. Available from: https://doi.org/10.1128/jcm.00204-19
- Neugent ML, Hulyalkar NV, Nguyen VH, Zimmern PE, De Nisco NJ. Advances in understanding the human urinary microbiome and its potential role in urinary tract infection. mBio. 2020;11(2):e00218-20. Available from: https://doi.org/10.1128/mbio.00218-20
- Moustafa A, Li W, Singh H, Moncera KJ, Torralba MG, Yu Y, et al. Microbial metagenome of urinary tract infection. Sci Rep. 2018;8(1):4333. Available from: https://doi.org/10.1038/s41598-018-22660-8
- Zhang Y, Wang H, Yan R, Wang K, Man J, Yang L. Research advances on the urinary microbiome in non-infectious urinary tract diseases: from community composition to clinical prospects. Front Cell Infect Microbiol. 2026;16:1728182. Available from: https://doi.org/10.3389/fcimb.2026.1728182
- Hashemi N, Tondro Anamag F, Javan Balegh Marand A, Rahnama'i MS, Herizchi Ghadim H, Salehi-Pourmehr H, et al. A systematic and comprehensive review of the role of microbiota in urinary chronic pelvic pain syndrome. Neurourol Urodyn. 2024;43(8):1859-1882. Available from: https://doi.org/10.1002/nau.25550
- Fu C, Zhang Y, Liang L, Lin H, Shan K, Liu F, et al. The microbiota in patients with interstitial cystitis/bladder pain syndrome: a systematic review. BJU Int. 2024;134(6):869-880. Available from: https://doi.org/10.1111/bju.16439
- Shoskes DA, Altemus J, Polackwich AS, Tucky B, Wang H, Eng C. The urinary microbiome differs significantly between patients with chronic prostatitis/chronic pelvic pain syndrome and controls as well as between patients with different clinical phenotypes. Urology. 2016;92:26-32. Available from: https://doi.org/10.1016/j.urology.2016.02.043
- Deng AQ, Yue SY, Niu D, Zhang DD, Hou BB, Zhang L, et al. The role of microbiota in the chronic prostatitis/chronic pelvic pain syndrome: a review. Front Microbiol. 2025;16:1488732. Available from: https://doi.org/10.3389/fmicb.2025.1488732
- Qin J, Shi X, Xu J, Yuan S, Zheng B, Zhang E, et al. Characterization of the genitourinary microbiome of 1,165 middle-aged and elderly healthy individuals. Front Microbiol. 2021;12:673969. Available from: https://doi.org/10.3389/fmicb.2021.673969
- Hrbacek J, Morais D, Cermak P, Hanacek V, Zachoval R. Alpha-diversity and microbial community structure of the male urinary microbiota depend on urine sampling method. Sci Rep. 2021;11(1):23758. Available from: https://doi.org/10.1038/s41598-021-03292-x
- Nickel JC, Stephens A, Landis JR, Chen J, Mullins C, van Bokhoven A, et al. Search for microorganisms in men with urologic chronic pelvic pain syndrome: a culture-independent analysis in the MAPP research network. J Urol. 2015;194(1):127-135. Available from: https://doi.org/10.1016/j.juro.2015.01.037
- Davasaz Tabrizi E, Sevil M, Arican E. Bioinformatic strategies in metagenomics of chronic prostatitis. World J Urol. 2025;43(1):188. Available from: https://doi.org/10.1007/s00345-025-05514-7
- Zheng Z, Hu J, Li W, Ma K, Zhang C, Li K, et al. Integrated microbiome and metabolome analysis reveals novel urinary microenvironmental signatures in interstitial cystitis/bladder pain syndrome patients. J Transl Med. 2023;21(1):266. Available from: https://doi.org/10.1186/s12967-023-04115-5
- Nickel JC, Stephens A, Landis JR, Mullins C, van Bokhoven A, Lucia MS, et al. Assessment of the lower urinary tract microbiota during symptom flare in women with urologic chronic pelvic pain syndrome: a MAPP network study. J Urol. 2016;195(2):356-362. Available from: https://doi.org/10.1016/j.juro.2015.09.075